Introduction
This page describes cloud-based research tools aligned with the HIMAT research program and the Ocean Observing Initiative.
OOI
HiMAT
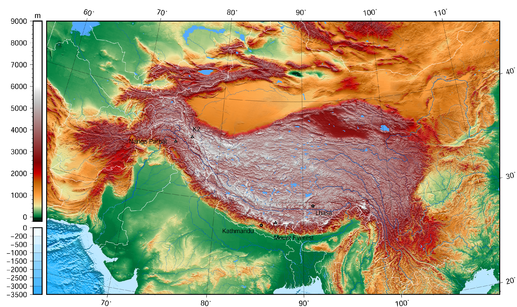
Project lead = Dr. Anthony Arendt. High Mountain Asia (HMA) extends from the Hindu Kush and Tien Shan in the west to the Eastern Himalaya. It is the world’s largest reservoir of perennial glaciers and snow outside the polar ice sheets. The region is home to a range of unique landforms, ecosystems, hazards, and cultures; and supplies water to more than a billion people. Changes in the region’s glaciers, snow, permafrost, and precipitation patterns have altered this water supply while also transforming regional ecology, land utilization practices, and the hazards associated with landslides and glacial-lake-outburst floods.
The goal of the HiMAT project is to use satellite remote sensing observations to characterize these changes, gain insight into the Earth system processes that control them, and inform decisions, management actions, and policy development.
Jupyter Notebook
First we suggest UW researchers sign on to the eScience mailing list. It’s the box on the bottom right-hand side of the front page of the escience website. You will receive notifications of upcoming events like software carpentry and data carpentry and cloud computing courses.
Our objective here is to pull raster data in NetCDF format from S3 to a Jupyter Notebook environment and proceed to explore those data. To do this requires several steps… and we are writing here both for the person configuring the environment and the person using the environment. Therefore we suggest bear in mind whether you are one, the other or both.
As the person building the environment you will secure resources on the cloud and configure JupyterHub. This acts like a dispensary of Jupyter Notebook environments for example for a research team or a set of students.
As the person using the environment you will have a sense of what programming language you prefer and what libraries you would like to have pre-installed. You also may wish to install additional libraries in your environment.
As the builder you’ll need a cloud account. We tend to talk about AWS, Azure and GCP.
- Get a cloud instance to work on
- Follow Zero To Jupyter Hub
- This guide is provided by the Berkeley Institute for Data Science (BIDS)
- The guide will help you set up authentication for your JupyterHub customers
- It helps you with administration and other follow-on tasks
- Have your customers start using the system both by building notebooks and by learning how to manage the environment
As the customer of the JupyterHub instance you will want to jump in head-first with both feet and try to start breaking things. This helps pave the way (once the broken things are fixed) for others to come in to the environment and not be hindered by things breaking. Here are some other notes on getting up to speed…
- To manage your environment you may need to open a terminal window from your Jupyter Notebook home page
- From here you can issue Linux commands and edit files, particularly batch files and script files
- You may need to use the vi editor… and the text may be very annoying colors
- Let’s fix that: Create a .vimrc file in your home directory (or append to existing) these three lines:
syntax off
set nohlsearch
set t_Co=0
Let’s go through the steps of installing a new Python package as the JupyterHub customer.
InSAR
- Got the AMI shared at me by Scott
- Assigned it to a c4.8xlarge (and it is Ubuntu)
- Attached 256GB of block storage
- Follow this to mount the drive…
% lsblk
% sudo mkdir /processing
% sudo mkfs -t ext4 /dev/xvdb
% sudo mount /dev/xvdb /processing
% sudo chown -R ubuntu /processing
These commands…
- give me the ID of the EBS as /dev/xvdb
- make a mount point called /processing
- make a file system of the correct type
- mount the storage
- change the ownership
This gets us close to processing tandem phase ERS data.
The tandem mission ran 21-Mar-95 through 5-Jun-96. There is orbital ephemeris available for this but unfortunately it is not public on the web. Rathre I used an aws command line interface command to grab all of the ephemeris from Scott.
% aws s3 sync s3://bucketname local-directory
After this it is a matter of configuring two .xml files with the correct pointers and running the insarApp.py program. Which is already in my path thanks to the AMI knowing.
After a bit of searching at https://vertex.daac.asf.alaska.edu we find a decent pair and add these to the queue. Order RAW format (L0) since L1 is amplitude only.
Everything below here is old
Objective and Approach
Solution
We have developed a database and web-based API to facilitate data sharing between the principal investigators. The tools were built on Amazon Web Services using managed services for the database (RDS) and elastic load balancing to manage and scale the website.
We also have geospatial data stored on a Geoserver (acs/acs_geoserver.html) with a Leaflet frontend to allow users to visualize the data.
Results
Related links
Status
- Recent activity
- Juneau meeting with Chinese Academy of Sciences: Completed with presentation decks recovered
- Work in progress
- Accumulating a comprehensive overview of the NASA projects and interrelationships
- Selected contributor datasets placed in cloud storage
- Jupyter Hub installed on the cloud
- Example scripts running on Jupyter Hub
- Idea: Progress towards comparative processes
- High-Level Objectives
- Work to characterize degree of certainty in HIMAT program results / outputs: “How sure are you?”
HiMAT data infrastructure
A core feature of the HiMAT project is the construction and utilization of data sharing tools to foster efficient collaboration, reproducible research and enhanced stakeholder engagement. Our cloud-based data infrastructure aims to addresses several of the challenges that often limit effective collaboration in such projects:
-
cross-team collaboration necessitates the sharing of preliminary data products which are not yet fully validated and may not be ready to share with the public. Existing data infrastructures primarily store completed datasets on public-facing servers. So there is a considerable gap in our provisioning of privately accessible, cloud-based computational tools for this kind of research.
-
datasets to be generated by HiMAT are particularly voluminous, for example high resolution satellite imagery. Many existing data centers are not set up to handle data of this size, and even if they are, it is unreasonable to be downloading datasets this large to local machines. Therefore we need methods to co-locate our processing/analysis with the location at which the data are stored.
-
the development collaborative tools calls for some degree of customization in our computational infrastructure if we are to integrate our products and provide decision support to the region. Therefore investigators need to have full access to both front and backend computational components, without the need to submit requests to third party agencies.
To address these challenges we are designing a multi-tiered approach to data handling:
